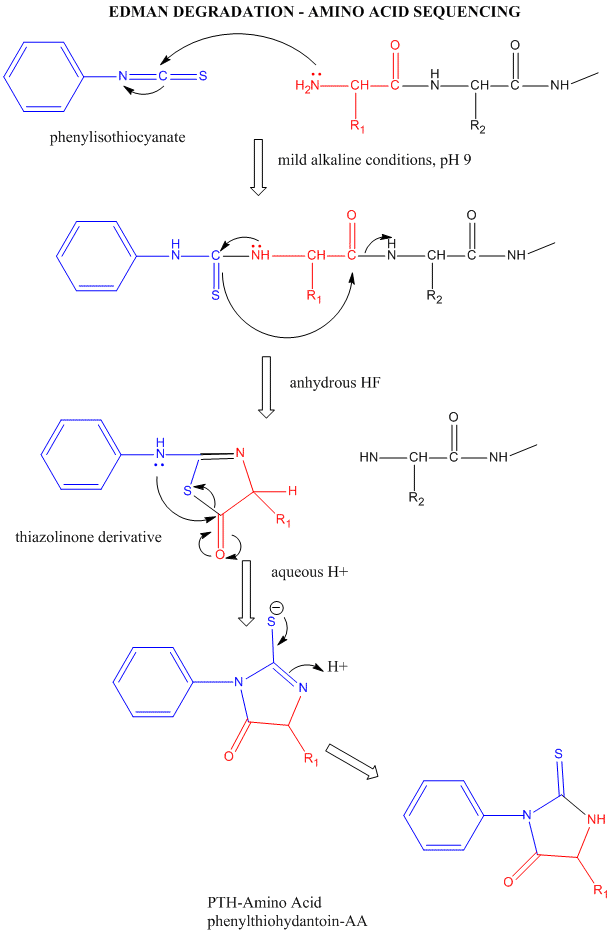
You may find it helpful to review the relationship between cyanates, isocyanates, thiocyanates and isothiocyanates.  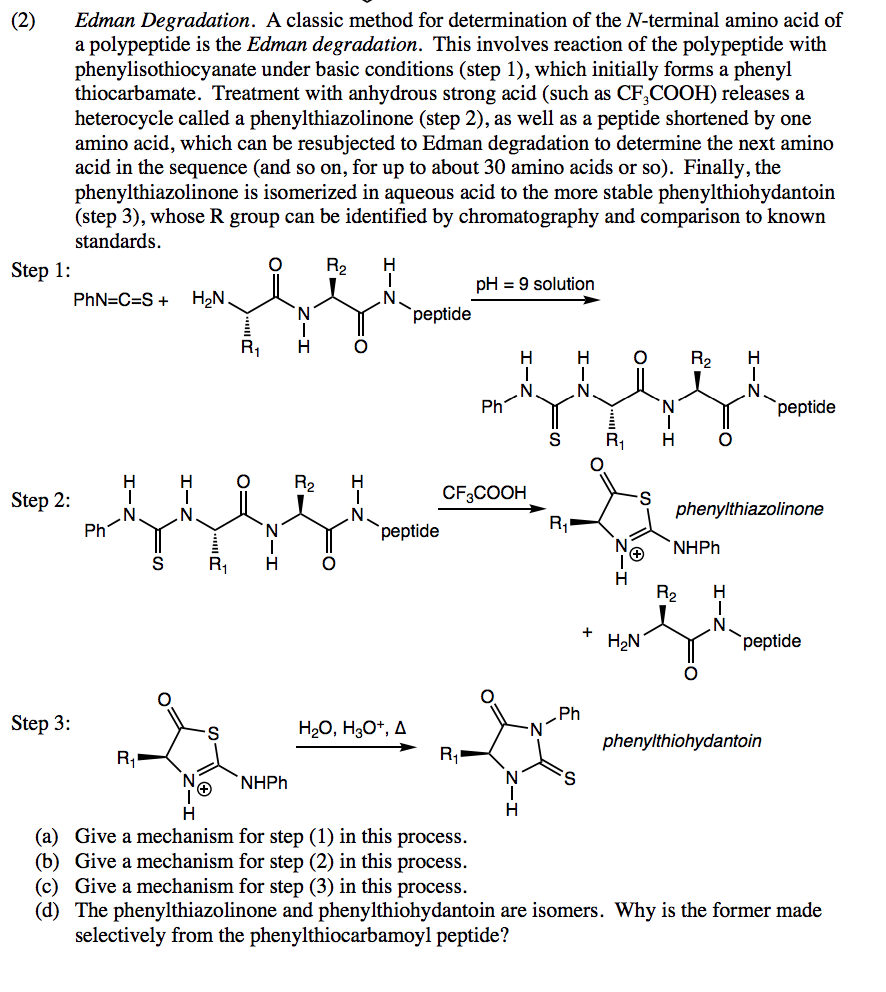
The reagent used in the Edman degradation is phenyl isothiocyanate. FIGURE 2: Mass spectrum representing the masses of peptides produced by the treatment of the heat shock protein tetradecapeptide with s-ETA (See question 25). The 2 very large peaks corresponding to the Edman reaction by-products dmptu and dptu are not included in the series of labels. These letters correspond to peak labels (from left to right). If you cannot read the labels placed on the top of each AA- PTH peaks on the profiles please refer to the series of letters placed on the right side of each panel. Panels a, b, and e represent the profile after 1.2, and 3 rounds of Edman degradation, respectively. The 2 very large peaks corresponding to the Edman reaction by products Ampla and dptu are not included in the series of labels These letters correspond to penk labels (from left to right). If you cannot read the labels placed on the top of each AA PTH pooks on the profiles please refer to the series of letters placed on the right side of each panel. Panels a, b, and represent the profile fer 1.2 and 3 rounds of Edman degradation, respectively. O) 24.00 200 16.00 12.00 8.00 a) DNSQTGEAHYRPMVFI Lee Lunchantin (b) 24.00 2000 16.00 باران b) DNSQTGEAHYRPMVFI 30.00 der 200 c) DNSTGEAHYRPMVFI FIGURE 1: HPLC profiles after Edman degradation (see question 1b). 3-b) 2 pts Provide an explanation for this observation using the mass difference between the original and the smaller peptide and from what you have learned in class about protein phosphorylation 3-e) 2 pts Suggest one method to confirm your hypothesis. 3-a) 2 pts Did you expect any cleavage with PFFA? Briefly explain. Question 3: The heat shock tetradecapeptide was also treated with PFFA, and 30% of the original peptide (Mass: 1642.8 Da) was converted into a smaller peptide (mass: 1562.8 Da). Note: One peak is produced by a post-translationally modified form of the uncleaved peptide. Identify the sequence of these 4 peptides using their measured mass and the table of masses that you got in one of your lecture handouts. 2-b) 4 pts The mass spectrum (Figure 2) shows that only 4 peaks were produced. 2-a) 4 pts Predict the sequence of all the cleaved peptides (assuming complete cleavage). Question 2: The heat shock protein tetradecapeptide has the following sequence: NH-CLNRQLSSGVSEIR-COOH This protein was treated with s-ETA and the cleavage fragments were analyzed by MALDI. Is your previous prediction confirmed by theses profiles? Briefly explain. 1-b) 4 pts The HPLC profiles (Figure 1) show the derivatized amino acid produced after round 1.2, and 3 of Edman degradation. Predict the sequence of all the peptides formed by chemical cleavage. Question 1: The a-melanocyte stimulating hormone is a short peptide whose sequence is: NH-SYSMEHFRWGKPV-COOH The peptide was submitted to chemical cleavage with s-ETA, and product of cleavage was sequenced by Edman degradation 1-a) 4pts. Recently, scientists have demonstrated that several chemical reagents can be used for the selective cleavage of peptide bonds in proteins: for example, S-ethyl trifluorothioacetate (s- ETA) selectively cleaves the peptide bond located on the amino-side of Ser and Thr residues, and similarly, pentafluoropropionic acid (PFPA) selectively cleaves the peptide bonds located on the carboxyl-side of Asp residues.
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
